- Multiplex Ligation dependent Probe Amplification (MLPA) is a multiplex PCR method used to detect abnormal copy numbers of up to 50 different genomic DNA or RNA sequences. Furthermore, MS-MLPA can detect DNA methylation changes.
- Is the most reliable and cost-effective method of detecting known deletion, duplications, and specific copy number variations (CNVs). Like array CGH, MLPA detects copy number changes and the interpretation of the results can be complicated by naturally occurring copy number variations. The probes or probe kits used have been selected and validated to reduce the likelihood of false positive or negative results.
MLPA – Multiplex ligation-dependent probe amplification

MLPA can be indicated across all life stages and is dependent on the condition in question and the clinical symptoms present. MLPA can be requested if:
- The patient is suspected to have a genetic disorder that could be caused by a deletion or duplication.
- Previous genetic tests were negative or only identified a single variant in a gene or condition that is associated with autosomal recessive inheritance.
MLPA can be used to diagnose specific disorders associated to:
- Deletions or duplications of specific regions, genes or exons
- Imprinting alterations .
- Uniparental disomy (UPD)
- Small rearrangements (small intragenic deletions)
MLPA can detect:
- Small deletions and rearrangements associated to specific regions, genes or exons
- Specific microdeletion syndromes
- Diseases caused by methylation defects (MS-MLPA)
- Specific Uniparental disomy
The most common genetic disorders detected by MLPA include:
- TSC1 deletion/duplication
- TSC2 deletion/duplication
- Congenital adrenal hyperplasia CYP21A2 (21-0H) deletion/duplication analysis
- ATP7B deletion/duplication
- HBB deletion/duplication
- HBA1 & HBA2 deletion/duplication
- IKBKG deletion/duplication analysis
- Ornithine transcarbamylase deficiency (OTC) deletion/duplication analysis
- Ataxia-telangiectasia (ATM) deletion/duplication
- Neurofibromatosis type 1 (NF1) deletion/duplication
- Neurofibromatosis type 2 (NF2) deletion/duplication
- Neurodegeneration with brain iron accumulation 2B (PLA2G6) deletion/duplication analysis
- Pantothenate kinase-associated degeneration (PANK2) deletion / duplication analysis
- Duchenne Muscular Dystrophy (DMD) deletion/duplication
- PMP22 deletion/duplication analysis
- Spinal Muscular Atrophy (SMN1/SMN2) deletion/duplication
- DiGeorge syndrome deletion/duplication analysis
- Cystic fibrosis (CFTR) gene deletion/duplication
- Prader-Willi/Angelman syndrome deletion/duplication
- Ad-hoc deletion/duplication analysis
There are two possible results that can be obtained from an MLPA test
-
Positive (Pathogenic and likely pathogenic)
A positive result indicates that a gene deletion or duplication has been identified in association with the disease phenotype under study. This scenario will allow to provide genetic counselling or personal guidance regarding possible medical treatments, disease progression, reproductive-/prevention-strategies and potential implications for other family members
-
Negative:
A negative result indicates that no disease-causing deletions or duplications were identified in the test performed. It does not guarantee that the induvial will be healthy or free from other genetic disorders or medical conditions. Additionally, a negative result does not rule out a genetic cause of the disease nor does it eliminate the risk for future offspring. However, if a negative test result is obtained and the variant in question is known to be present in affected family members, this then rules out a diagnosis of that genetic disorder in the proband. A negative result may be explained by several causes, including limited genetic knowledge and limitations associated to the used methodology
Sample Requirements
For genetic testing through next generation sequencing, the following sample types are accepted. A thorough labelling of the tube with unique identifying information is suggested, incorrect labelling can lead to rejection of the sample. The minimum required information to identify and accept a sample is – Patient’s full name, Date of birth, Gender and Medical Record Number.
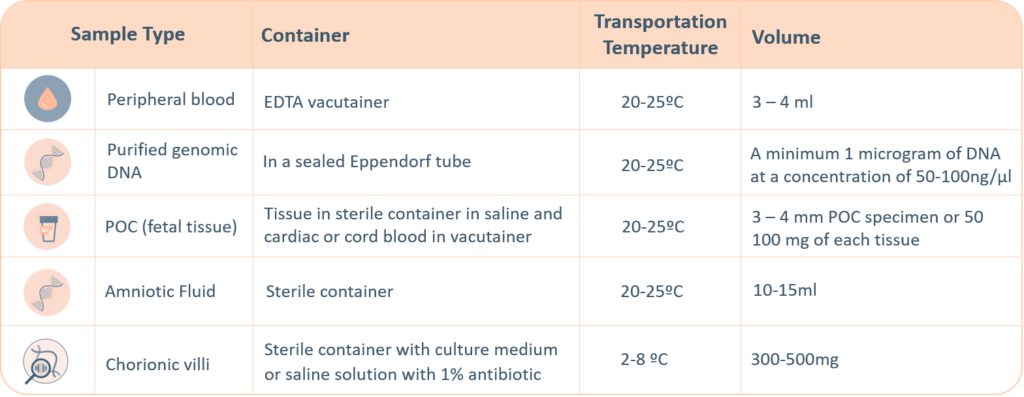
- Maternal blood sample must be sent with all products of conception, CVS and Amnio samples
- Precedence will be given to all prenatal samples
The ‘informed consent’ form and the ‘test requisition from’ (included within the provided kit) must be properly filled-in and signed by the patient and sent with the samples inside the shipping box or by e-mail to the laboratory. Igenomix will send you all the documents needed for the pick-up and transportation of the appropriate kit to our laboratory
Methodology
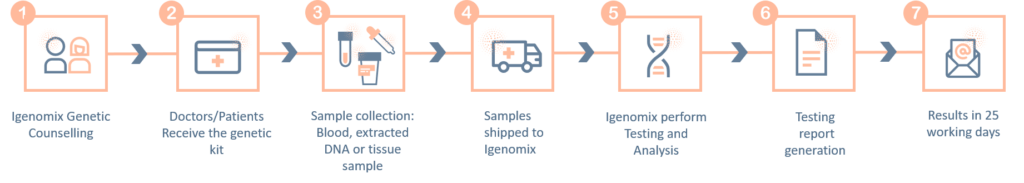
Limitations
MLPA cannot detect
- Balanced chromosomal rearrangements
- Telemoric deletions and duplications
- Deletions and duplications that are not identified by the MLPA probes used
- Point mutations, small insertions and deletions
- Sequence repeats or disorders caused by mutations in mitocondrial DNA
Multiplex Ligation-dependent Probe Amplification. MLPA probe sets and reagents from MRC-Holland were used for copy number analysis of specific targets versus known control samples. False positive or negative results may occur due to rare sequence variants in target regions detected by MLPA probes. Analytical sensitivity and specificity of the MLPA method are both 99%. MLPA cannot detect any changes that lie outside the target sequence of the probes and will not detect most inversions or translocations. Even when MLPA did not detect any aberrations, the possibility remains that biological changes in that gene or chromosomal region do exist but remain undetected. The MLPA test will not detect the point mutations in the DMD genes. A point mutation or polymorphism in the sequence detected by a probe, which results in reduced probe binding efficiency, can also cause a reduction in relative peak area. Therefore, single exon deletions detected by MLPA should always be confirmed by other methods like multiplex PCR or sequencing.
Why Choose IGENOMIX?
High-Quality interpretation
Genetic counselling E-learning platform
Easy & Fast Online tests management
Local Labs over the Middle East
All Technical capabilities In-House
100% In-House sequencing and Bio IT
CAP & ISO accredited lab
Using international, population and
diseases Databases, including HGMD





